SLIDE 1 Chapter 3: Density dependence
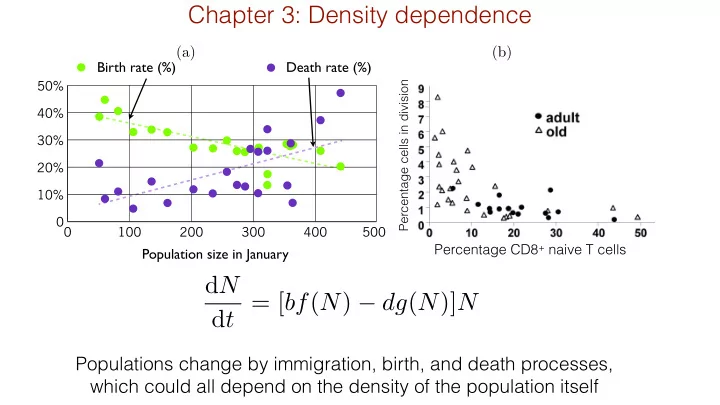
(a) (b)
Percentage CD8+ naive T cells
Percentage cells in division
Populations change by immigration, birth, and death processes, which could all depend on the density of the population itself
dN dt = [bf(N) − dg(N)]N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 2 Population density Population density Per capita birth or death rate (a) (b)
death rate death rate birth rate birth rate steady state steady state
dN dt > 0 dN dt > 0 dN dt < 0 dN dt < 0
dN dt = h b − d ⇣ 1 + N k ⌘i N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
d b b d
¯ N = k b − d d = k(R0 − 1)
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
dN dt = [b − d f(N)]N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
Population density
F(N)
F(N) = d + cN = d f(N) ↔ f(N) = 1 + N/k
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 3 Population density
total death rate production rate steady state
dN dt > 0 dN dt < 0
Production or death rate
dN dt = s − d ⇣ 1 + N k ⌘ N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
¯ N = −dk ± p dk(dk + 4s) 2d
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
s

SLIDE 4 Population density Population density Per capita birth or death rate (a) (b)
death rate death rate birth rate birth rate steady state steady state
dN dt > 0 dN dt > 0 dN dt < 0 dN dt < 0
d b b d
dN dt = [bf(N) − d]N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
¯ N = k ⇣ 1 − d b ⌘ = k ⇣ 1 − 1 R0 ⌘
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
dN dt = h b ⇣ 1 − N k ⌘ − d i N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
Population density
F(N)
F(N) = b − cN = bf(N) ↔ f(N) = 1 − N/k
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 5 dN dt = s ⇣ 1 − N k ⌘ − dN
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
Population density
total death rate production rate steady state
dN dt > 0 dN dt < 0
Production or death rate
¯ N = sk dk + s
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
s

SLIDE 6
(a)
Time N(t) K
(b)
N r(1 N/K) r K
(c)
N r(1 (N/K)m) K
Logistic growth: dN
dt = rN(1 N/K) , with solution N(t) = KN(0) N(0) + e−rt(K N(0))
m=0.5 m=2

SLIDE 7 (a)
Time N(t) K
(b)
N r(1 N/K) r K
(c)
N r(1 (N/K)m) K
Generalized logistic growth:
m=0.5 m=2
dN dt = rN(1 − (N/K)m) , with N(t) = K [1 − (1 − [K/N(0)]m)e−rmt]1/m
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 8
Human logistic growth
Human population in Monroe Country, West Virginia

SLIDE 9 (a)
Time N(t)
(b)
Time N(t)
dN dt = s ⇣ 1 − N k ⌘ − dN
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
dN dt = s − d ⇣ 1 + N k ⌘ N
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 10 Density dependent birth
Corn Salicornia Grizzly bear
10 100 100 1,000 10,000 Seeds planted per m2 Average number of seeds per reproducing individual (a) 10 20 30 40 Density of females 50 60 70 80 4.0 3.8 3.6 3.4 3.2 3.0 2.8 Clutch size Plantain (b) Song sparrow

SLIDE 11 Non-linear density dependence
f(x) = max(0, 1 − [x/k]n)
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
f(x) = min(1, [x/k]n)
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
f(x) = xn hn + xn
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
g(x) = 1 1 + (x/h)n
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
f(x) = 1 − e− ln[2]x/h
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
g(x) = e− ln[2]x/h
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>

SLIDE 12 dN dt = (bf(N) − d)N ,
(a)
N f(N) k b d + −
(b)
N f(N) b k d + −
(c)
N f(N) b k d + −
f(N) = 1 1 + N/k , f(N) = 1 1 + [N/k]2 and f(N) = e− ln[2]N/k

SLIDE 13 dN dt = (bf(N) − d)N ,
(a)
N f(N) k b d + −
(b)
N f(N) b k d + −
(c)
N f(N) b k d + −
f(N) = 1 1 + N/k , f(N) = 1 1 + [N/k]2 and f(N) = e− ln[2]N/k
Function f(0) f(k) f(∞) R0 Carrying capacity Eq. f(N) = max(0, 1 − [N/k]m) 1 b/d ¯ N = k m p 1 − 1/R0 (3.12) f(N) = 1/(1 + N/k) 1 0.5 b/d ¯ N = k(R0 − 1) (3.14) f(N) = 1/(1 + [N/k]2) 1 0.5 b/d ¯ N = k√R0 − 1 (3.14) f(N) = e− ln[2]N/k 1 0.5 b/d ¯ N = (k/ ln[2]) ln[R0] (3.14)

SLIDE 14 Density dependent death
(a)
N d + δf(N) d d + δ
(b)
N d[1 + (N/k)m] d
dN dt = (b − d[1 + (N/k)m])N
dN dt = [b − d − δf(N)]N ,
+

SLIDE 15 Positive density dependence
(a)
Time N(t) N(0) K
(b)
N bf(N)g(N)
dN dt = (bf(N)g(N) d)N
dN dt = (bg(N) d[1 + (N/k)m])N ,

SLIDE 16
Regression to the mean
n <- 100; data <- rnorm(n,1,0.1);hist(data) N <- data[1:(n-1)]; r <- (data[2:n]-N)/N plot(N,r,type="p") lm(r~N,as.data.frame(cbind(N,r)))

SLIDE 17
Smith-Martin model (first ignoring death):
is dN/dt = (p − d)N, w Cell division takes time
Conventional ODE:
dA(t) dt = 2pAt−∆ − pA(t) and dB(t) dt = pA(t) − pAt−∆
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
dA(t) dt = 2pAt−∆e−d∆−(p+d)A(t) and dB(t) dt = pA(t)−dB(t)−pAt−∆e−d∆
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
Smith-Martin model with death:

SLIDE 18
dA dt = 2n ∆ Bn−(p+d)A , dB1 dt = pA− ⇣ d+ n ∆ ⌘ B1 and dBi dt = n ∆ (Bi−1−Bi)−dBi
Time delays implemented as many small steps
Smooth the time delay by many (n) small steps:
dA(t) dt = 2pAt−∆e−d∆−(p+d)A(t) and dB(t) dt = pA(t)−dB(t)−pAt−∆e−d∆
<latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit><latexit sha1_base64="(nul)">(nul)</latexit>
Smith-Martin model with death: