Mary K. Campbell Shawn O. Farrell
- Chapter Twelve
Protein Synthesis: Translation of the Genetic Message
Paul D. Adams • University of Arkansas
1
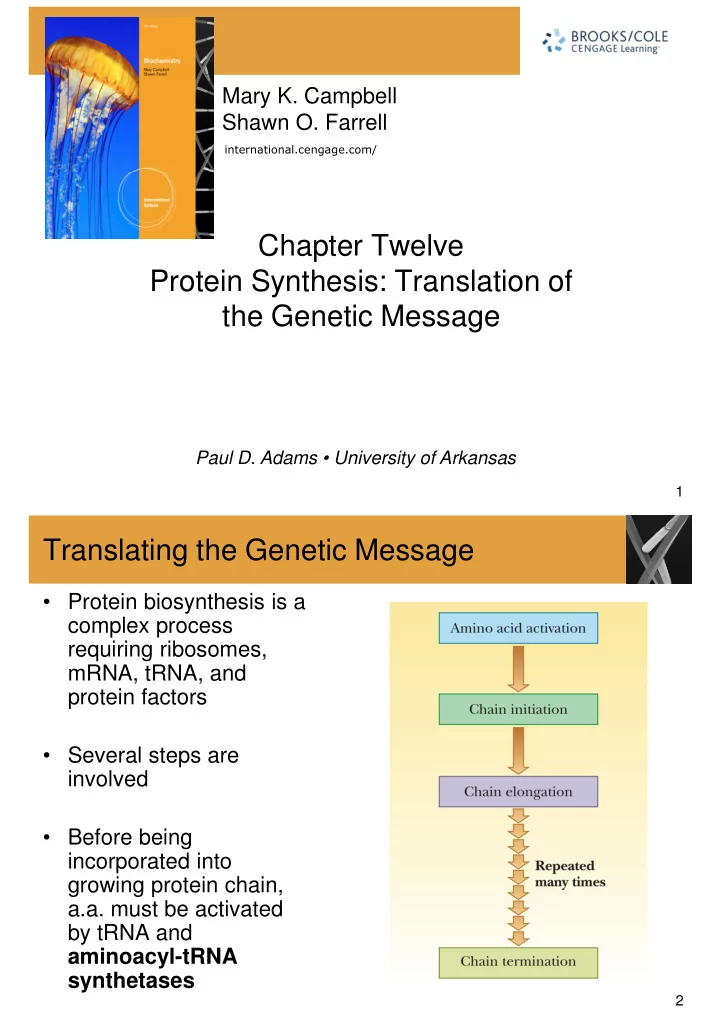
Translating the Genetic Message
- Protein biosynthesis is a
complex process requiring ribosomes, mRNA, tRNA, and mRNA, tRNA, and protein factors
- Several steps are
involved
- Before being
- Before being
incorporated into growing protein chain, a.a. must be activated by tRNA and aminoacyl-tRNA synthetases
2