SLIDE 1
Review questions BSC3052 FST and PVA
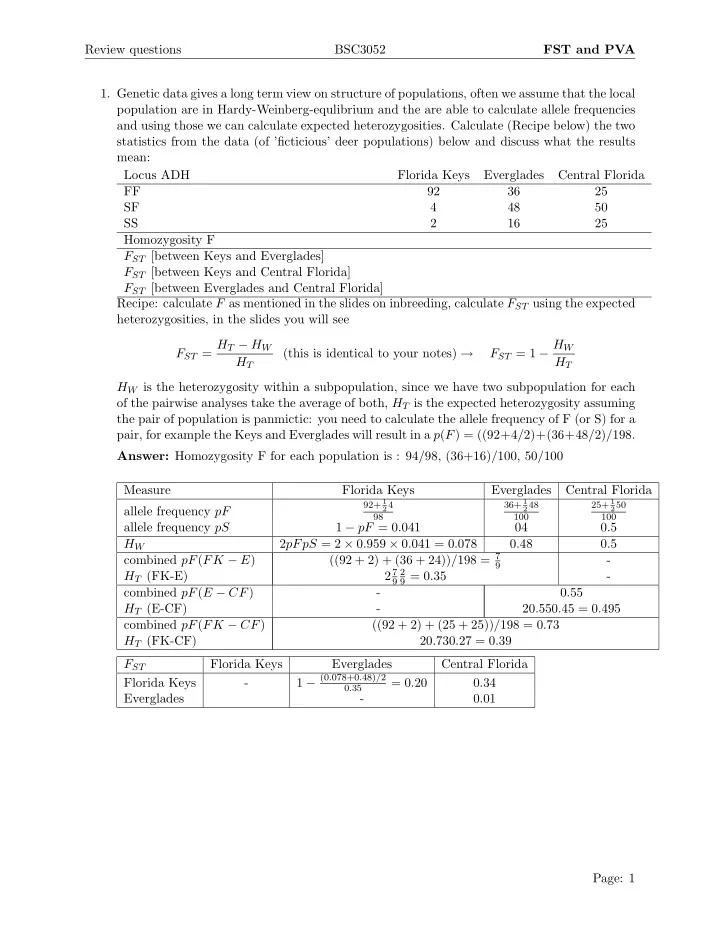
- 1. Genetic data gives a long term view on structure of populations, often we assume that the local
population are in Hardy-Weinberg-equlibrium and the are able to calculate allele frequencies and using those we can calculate expected heterozygosities. Calculate (Recipe below) the two statistics from the data (of ’ficticious’ deer populations) below and discuss what the results mean: Locus ADH Florida Keys Everglades Central Florida FF 92 36 25 SF 4 48 50 SS 2 16 25 Homozygosity F FST [between Keys and Everglades] FST [between Keys and Central Florida] FST [between Everglades and Central Florida] Recipe: calculate F as mentioned in the slides on inbreeding, calculate FST using the expected heterozygosities, in the slides you will see FST = HT − HW HT (this is identical to your notes) → FST = 1 − HW HT HW is the heterozygosity within a subpopulation, since we have two subpopulation for each
- f the pairwise analyses take the average of both, HT is the expected heterozygosity assuming
the pair of population is panmictic: you need to calculate the allele frequency of F (or S) for a pair, for example the Keys and Everglades will result in a p(F) = ((92+4/2)+(36+48/2)/198. Answer: Homozygosity F for each population is : 94/98, (36+16)/100, 50/100 Measure Florida Keys Everglades Central Florida allele frequency pF
92+ 1
24
98 36+ 1
2 48
100 25+ 1
250
100
allele frequency pS 1 − pF = 0.041 04 0.5 HW 2pFpS = 2 × 0.959 × 0.041 = 0.078 0.48 0.5 combined pF(FK − E) ((92 + 2) + (36 + 24))/198 = 7
9
- HT (FK-E)
27
9 2 9 = 0.35
- combined pF(E − CF)
- 0.55
HT (E-CF)
- 20.550.45 = 0.495
combined pF(FK − CF) ((92 + 2) + (25 + 25))/198 = 0.73 HT (FK-CF) 20.730.27 = 0.39 FST Florida Keys Everglades Central Florida Florida Keys
- 1 − (0.078+0.48)/2
0.35
= 0.20 0.34 Everglades
- 0.01